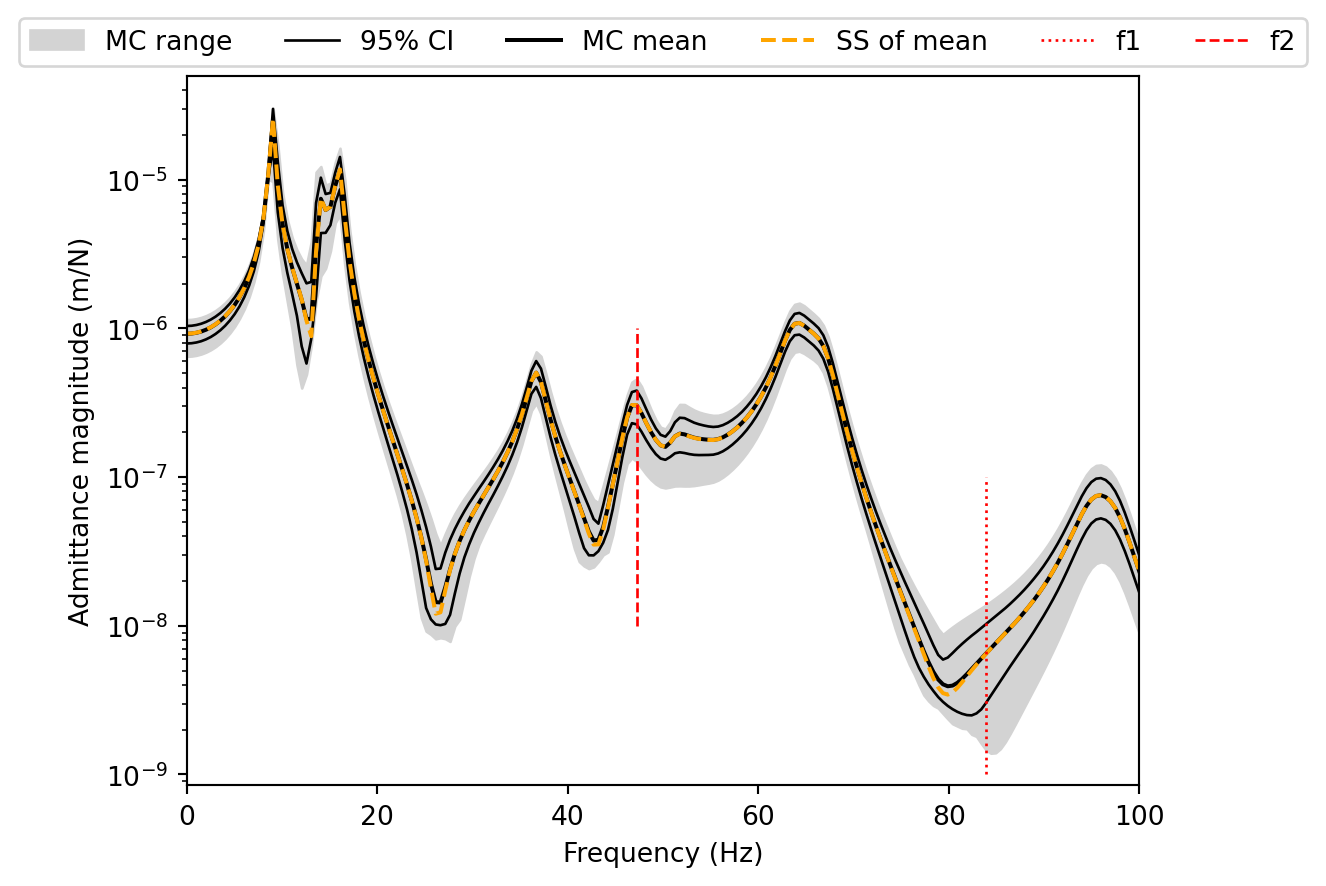
###### Frequency 1 ########
μY_real = np.mean(y0[:,f1])
μY_imag = np.mean(y1[:,f1])
ΣY = np.cov(np.stack((y0[:,f1], y1[:,f1])))
import matplotlib.gridspec as gridspec
# Generate plot
# fig = plt.figure(figsize=(5, 5))
fig = plt.figure(figsize=(9, 4))
outer = gridspec.GridSpec(1, 2, wspace=0.3)
gs1 = gridspec.GridSpecFromSubplotSpec(2, 2, width_ratios=(4, 1), height_ratios=(1, 4), subplot_spec=outer[0], wspace=0.1, hspace=0.1)
ax = fig.add_subplot(gs1[1, 0])
ax_histx = fig.add_subplot(gs1[0, 0], sharex=ax)
ax_histy = fig.add_subplot(gs1[1, 1], sharey=ax)
ax_histy.set_xticks([])
ax_histx.set_yticks([])
scatter_hist(y0[:,f1], y1[:,f1], ax, ax_histx, ax_histy, 50)
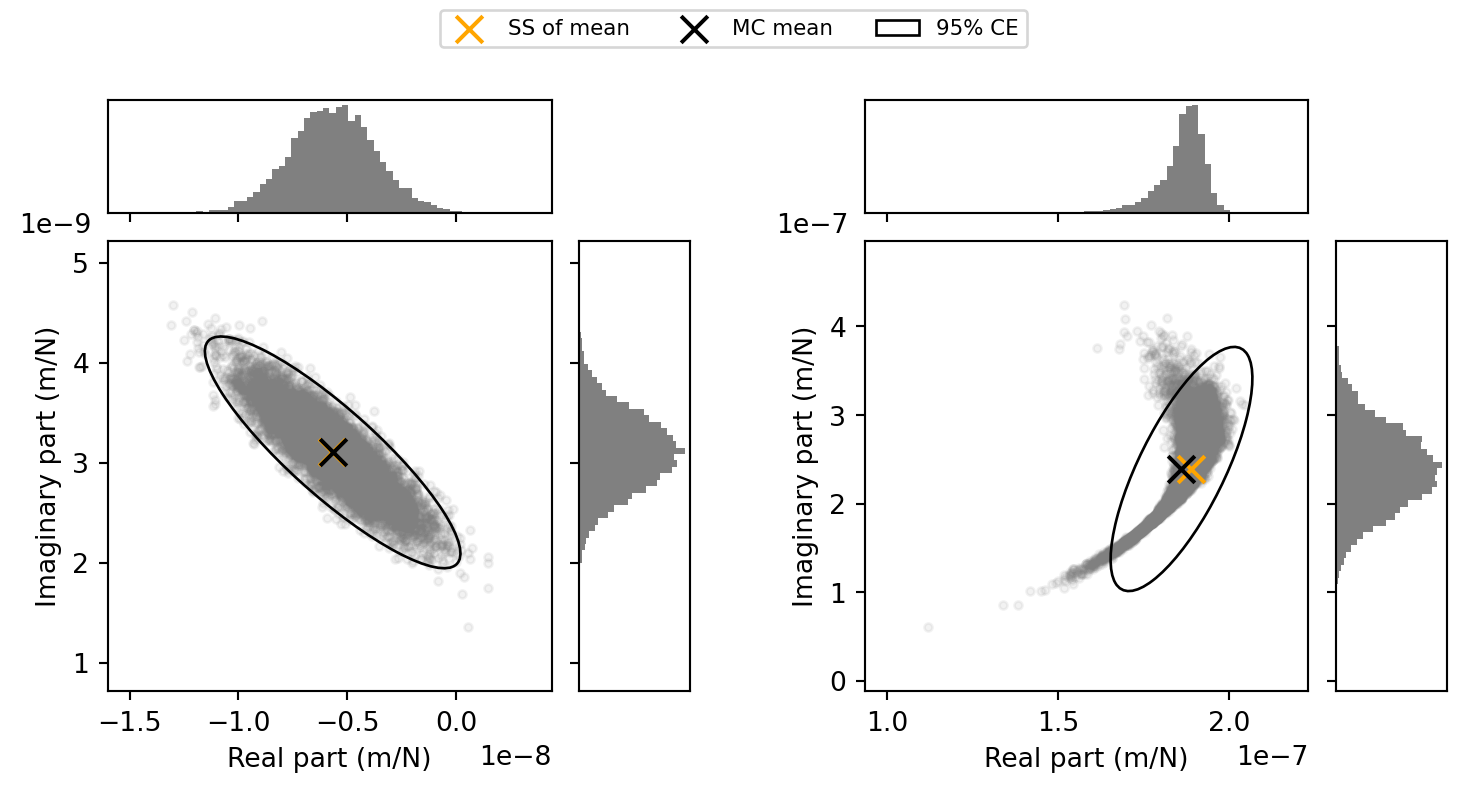
ax.scatter(np.real(Y_C_dual_μ[f1, 0, -1]), np.imag(Y_C_dual_μ[f1, 0, -1]), s=100,marker='x',color='orange',label='SS of mean')
ax.scatter(μY_real, μY_imag, s=100, marker='x',color='black',label='MC mean')
ax.set_xlabel('Real part (m/N)')
ax.set_ylabel('Imaginary part (m/N)')
Δx = 0.2
Δy = 0.2
ax.set_xlim(np.min(y0[:,f1])-Δx*np.abs(np.max(y0[:,f1])-np.min(y0[:,f1])), np.max(y0[:,f1])+Δx*np.abs(np.max(y0[:,f1])-np.min(y0[:,f1])))
ax.set_ylim(np.min(y1[:,f1])-Δy*np.abs(np.max(y1[:,f1])-np.min(y1[:,f1])), np.max(y1[:,f1])+Δy*np.abs(np.max(y1[:,f1])-np.min(y1[:,f1])))
ax.ticklabel_format(axis='both',style='scientific',scilimits=(0,0))
t = ax.yaxis.get_offset_text()
t.set_x(-0.2)
ell1 = confidence_ellipse([μY_real, μY_imag], ΣY, ax, n_std=3.0, edgecolor='black', linestyle='solid',label='95% CE')
###### Frequency 2 ########
μY_real = np.mean(y0[:,f2])
μY_imag = np.mean(y1[:,f2])
ΣY = np.cov(np.stack((y0[:,f2], y1[:,f2])))
# Generate plot
gs2 = gridspec.GridSpecFromSubplotSpec(2, 2, width_ratios=(4, 1), height_ratios=(1, 4), subplot_spec=outer[1], wspace=0.1, hspace=0.1)
ax = fig.add_subplot(gs2[1, 0])
ax_histx = fig.add_subplot(gs2[0, 0], sharex=ax)
ax_histy = fig.add_subplot(gs2[1, 1], sharey=ax)
ax_histy.set_xticks([])
ax_histx.set_yticks([])
scatter_hist(y0[:,f2], y1[:,f2], ax, ax_histx, ax_histy, 50)
ax.scatter(np.real(Y_C_dual_μ[f2, 0, -1]), np.imag(Y_C_dual_μ[f2, 0, -1]), s=100,marker='x',color='orange',label='SS of mean')
ax.scatter(μY_real, μY_imag, s=100, marker='x',color='black',label='MC mean')
ax.set_xlabel('Real part (m/N)')
ax.set_ylabel('Imaginary part (m/N)')
Δx = 0.2
Δy = 0.2
ax.set_xlim(np.min(y0[:,f2])-Δx*np.abs(np.max(y0[:,f2])-np.min(y0[:,f2])), np.max(y0[:,f2])+Δx*np.abs(np.max(y0[:,f2])-np.min(y0[:,f2])))
ax.set_ylim(np.min(y1[:,f2])-Δy*np.abs(np.max(y1[:,f2])-np.min(y1[:,f2])), np.max(y1[:,f2])+Δy*np.abs(np.max(y1[:,f2])-np.min(y1[:,f2])))
ax.ticklabel_format(axis='both',style='scientific',scilimits=(0,0))
t = ax.yaxis.get_offset_text()
t.set_x(-0.2)
ell1 = confidence_ellipse([μY_real, μY_imag], ΣY, ax, n_std=3.0, edgecolor='black', linestyle='solid',label='95% CE')
handles, labels = ax.get_legend_handles_labels()
fig.legend(handles, labels, loc='upper center',ncol=3,fontsize=8)
plt.show()