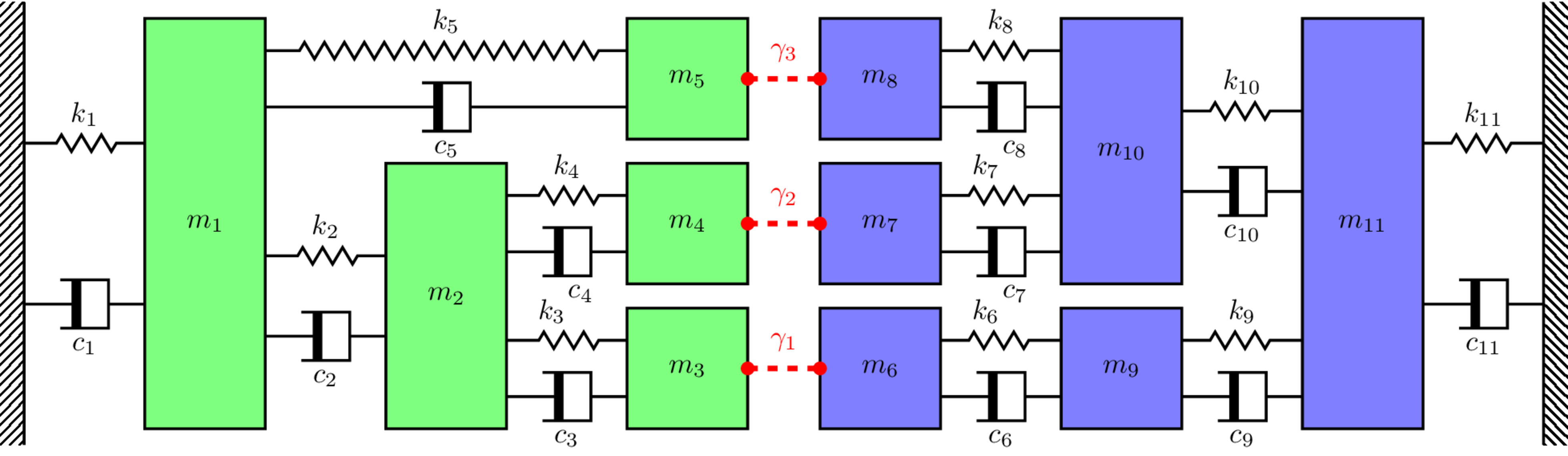
ηj = 0.5 # Loss factor of coupling element
kj = 10000*(1+1j*ηj) # Stiffness of coupling element
# Redefine mass and stiffness matrices for resiliently coupled assembly
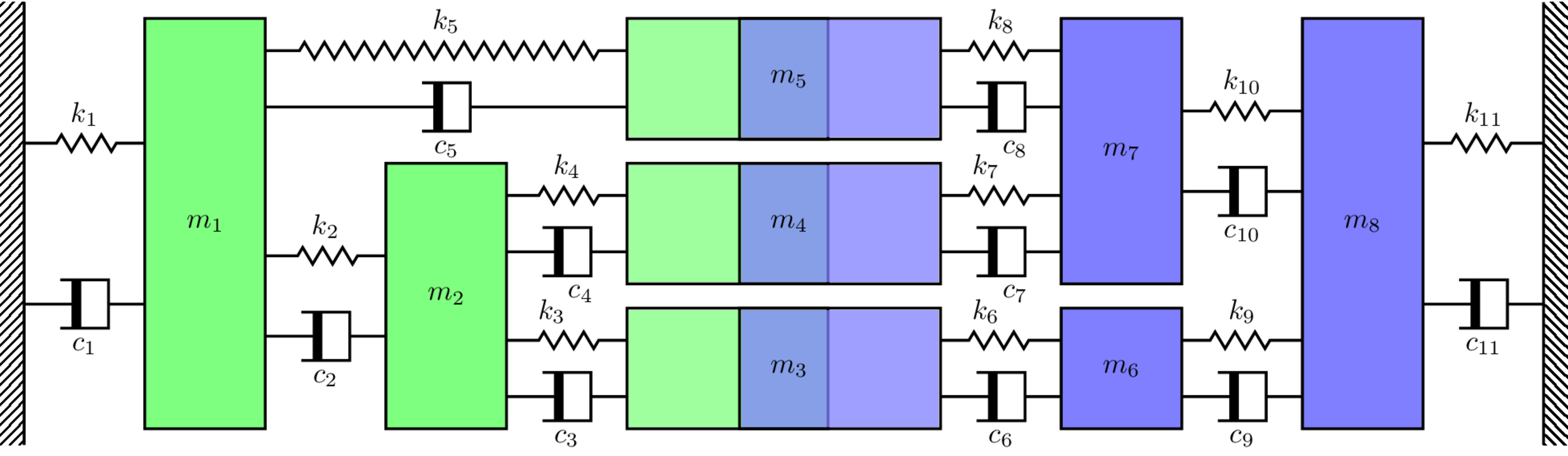
M_C_res = sp.linalg.block_diag(M_A,M_B)
K_C_res = np.array([[k1+k2+k5, -k2, 0, 0, -k5, 0, 0, 0, 0, 0, 0],
[-k2, k2+k3+k4, -k3, -k4, 0, 0, 0, 0, 0, 0, 0],
[0, -k3, k3+kj, 0, 0, -kj, 0, 0, 0, 0, 0],
[0, -k4, 0, k4+kj, 0, 0, -kj, 0, 0, 0, 0],
[-k5, 0, 0, 0, k5+kj, 0, 0, -kj, 0, 0, 0],
[0, 0, -kj, 0, 0, k6+kj, 0, 0, -k6, 0, 0],
[0, 0, 0, -kj, 0, 0, k7+kj, 0, 0, -k7, 0],
[0, 0, 0, 0, -kj, 0, 0, k8+kj, 0, -k8, 0],
[0, 0, 0, 0, 0, -k6, 0, 0, k6+k9, 0, -k9],
[0, 0, 0, 0, 0, 0, -k7, -k8, 0, k7+k8+k10, -k10],
[0, 0, 0, 0, 0, 0, 0, 0, -k9, -k10, k9+k10+k11]])
Y_C_res = np.linalg.inv(K_C_res - np.einsum('i,jk->ijk',ω**2,M_C_res)) # Compute coupled FRF directly
# For primal formulation:
Z_j = np.array([[kj, -kj],[-kj, kj]]) # Dynamic stiffness matrix of flexible connection
# New Boolean equilibrium matrix:
L = np.array([[1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[0,0,1,0,0,1,0,0,0,0,0,0,0,0,0,0,0],
[0,0,0,1,0,0,0,1,0,0,0,0,0,0,0,0,0],
[0,0,0,0,1,0,0,0,0,1,0,0,0,0,0,0,0],
[0,0,0,0,0,0,1,0,0,0,0,1,0,0,0,0,0],
[0,0,0,0,0,0,0,0,1,0,0,0,1,0,0,0,0],
[0,0,0,0,0,0,0,0,0,0,1,0,0,1,0,0,0],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1]]).T
print("The Boolean coupling matrix L.T:")
print(L.T)
print("\nThe Boolean coupling matrix B:")
print(B)
# For dual formulation:
γ = 1/kj # Joint flexibility (inverse of joint point stiffness)
Γ = np.diag([γ, γ, γ]) # Flexibility matrix for all joints (diagonal because no cross-coupling)
# Allocate some space
Y_C_primal_res = np.zeros_like(Y)
Y_C_dual_res = np.zeros_like(Y)
for fi in range(f.shape[0]): # Loop over frequency and compute coupled FRFs
Z = sp.linalg.block_diag(np.linalg.inv(Y_A[fi,:,:]), Z_j, Z_j, Z_j,np.linalg.inv(Y_B[fi,:,:])) # Block diagonal impedance matrix for primal method
Y_C_primal_res[fi,:,:] = np.linalg.inv(L.T@Z@L)
Y_C_dual_res[fi,:,:] = Y[fi,:,:] - Y[fi,:,:]@B.T@np.linalg.inv(B@Y[fi,:,:]@B.T + Γ)@B@Y[fi,:,:]
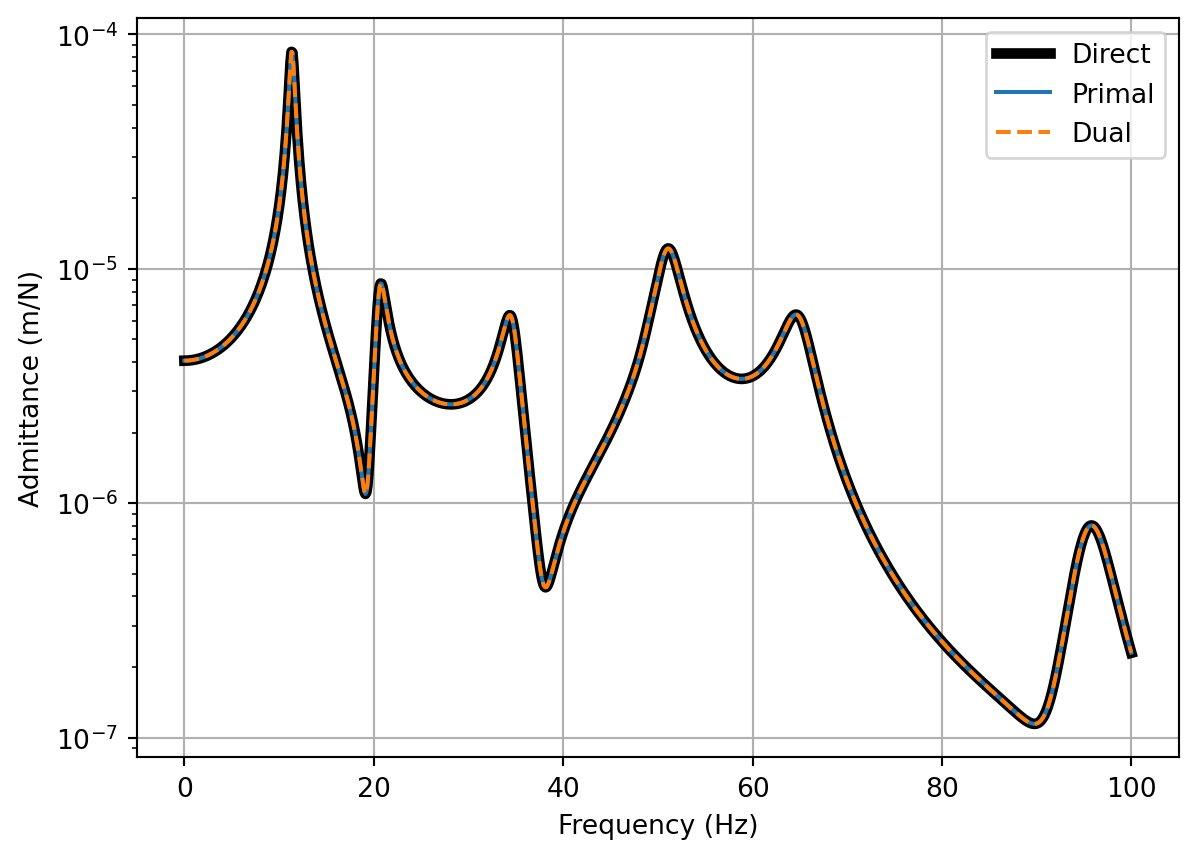
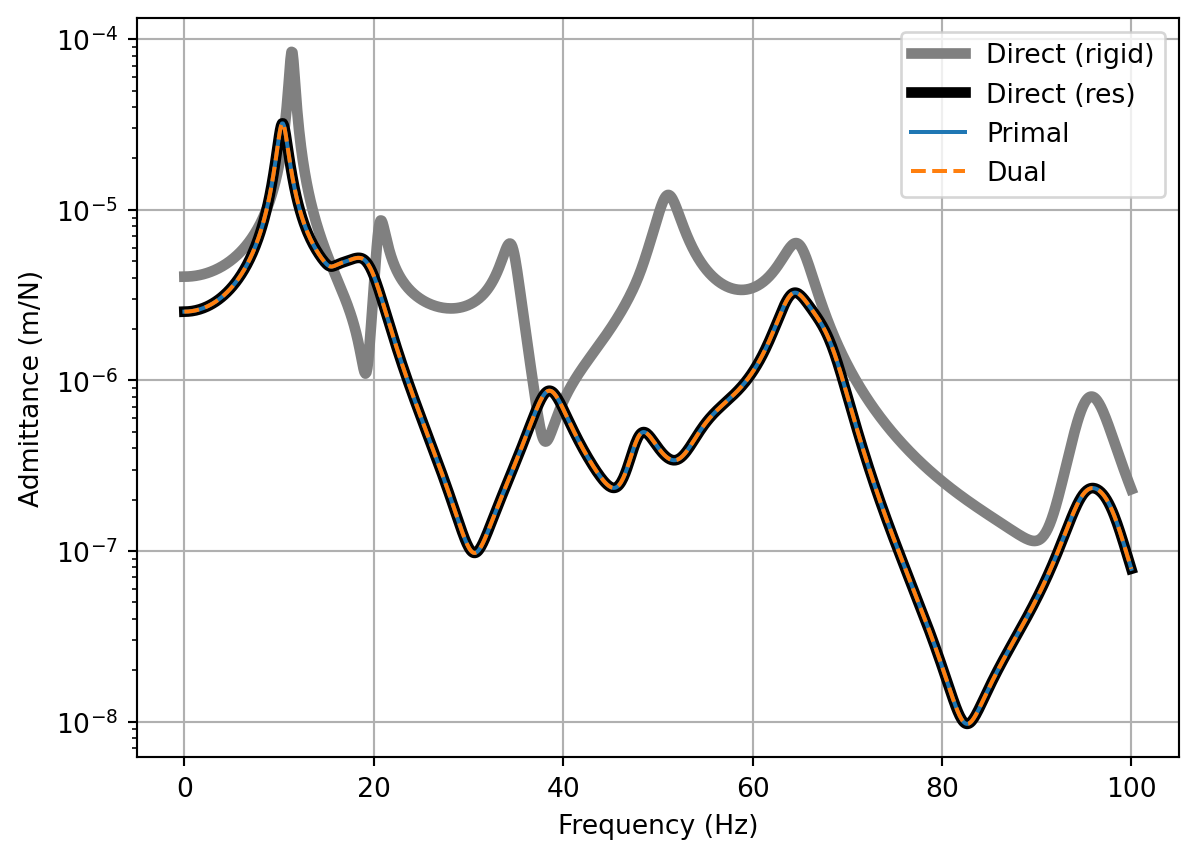
fig, ax = plt.subplots()
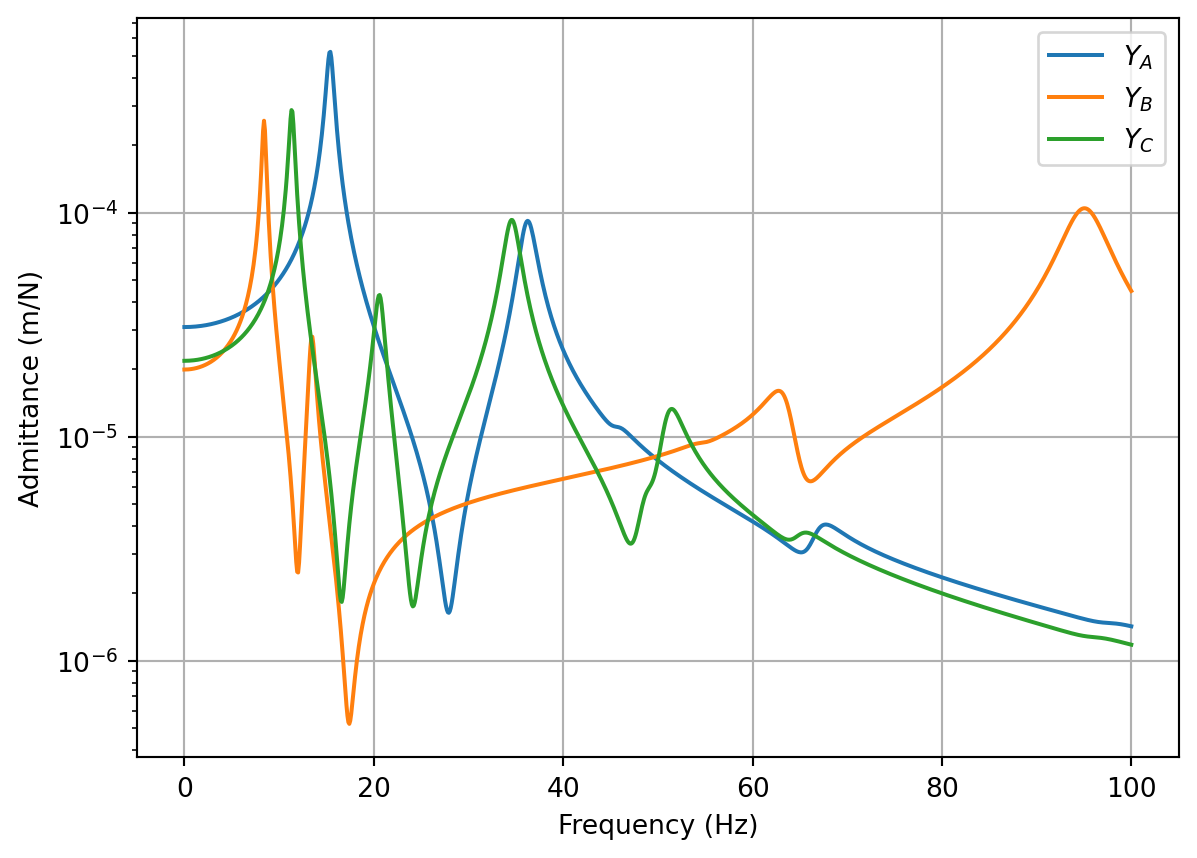
ax.semilogy(f, np.abs(Y_C[:,0,-1]),color='gray',label='Direct (rigid)',linewidth=4)
ax.semilogy(f, np.abs(Y_C_res[:,0,-1]),color='black',label='Direct (res)',linewidth=4)
ax.semilogy(f, np.abs(Y_C_primal_res[:,0,-1]),label='Primal')
ax.semilogy(f, np.abs(Y_C_dual_res[:,0,-1]),label='Dual',linestyle='dashed')
ax.grid(True)
ax.set_xlabel('Frequency (Hz)')
ax.set_ylabel('Admittance (m/N)')
plt.legend()
plt.show()